Contrast Sensitivity: Lesson 2
In this lesson, you will learn how to:
- duplicate an experiment,
- set experimental conditions through the Conditions method,
- move the events using drag and drop,
- and fit a model to measurements.
This lesson assumes you have been through Lesson 1 of this tutorial and become familiar with adding and inspecting the various types of events.
EFFECT OF SPATIAL FREQUENCY
In Lesson 1, you learned how to create a basic experiment to measure contrast threshold for a single spatial frequency. This lesson makes the basic contrast sensitivity task more elaborate by measuring the effect of the Gabor stimuli's spatial frequency. The measurements will be made through interleaved staircase methods, each of them specifying one experimental condition (spatial frequency).
Step 1: Duplicating the Experiment
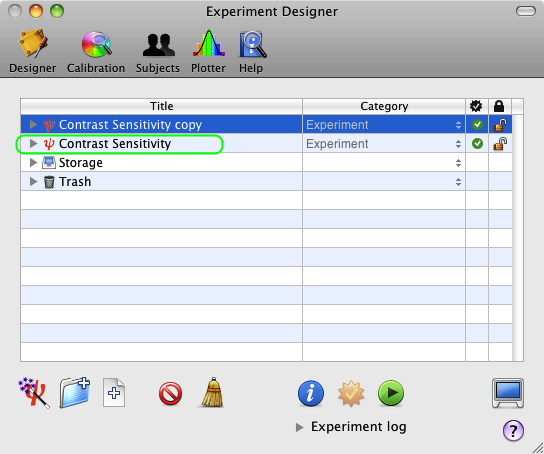
From the Designer panel, select the Contrast Sensitivity experiment designed in Lesson 1, and press the Apple-D keystroke (or select Duplicate from the Edit menu). The duplicate appears at the top level of the table with the 'copy' suffix added to the name of the original event. Rename the copy as Contrast Sensitivity 2.
Tip: There are alternative ways to duplicate an event:
- Using the regular Copy/Paste with the Apple-C & Apple-V keystrokes; events copied this way are always pasted in the Storage area of the table.
- Option-drag an event to make a copy at the dragged position.
Step 2: Adding and Customizing a "Conditions" Event
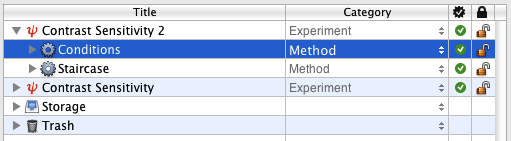
Select the Contrast Sensitivity 2 experiment, reveal its hierarchy, and insert a new group event using the '+' folder icon. Set the name and category of this new event to Conditions and Method, respectively.
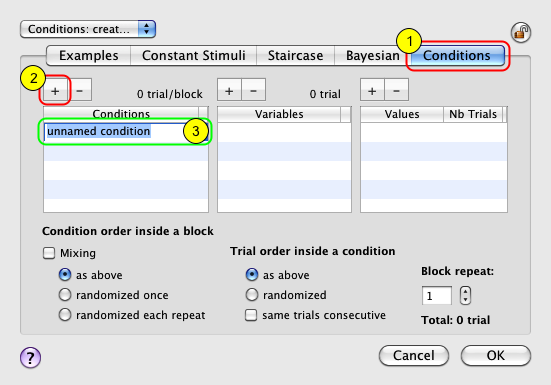
Select the new event named Conditions, and click on the Inspector button to inspect and modify its properties:
1) Click on the Conditions tab to select the conditions builder.
2) Click on the '+' button to add a new condition (called unnamed condition by default).
3) Rename it as Effect of Spatial Frequency.
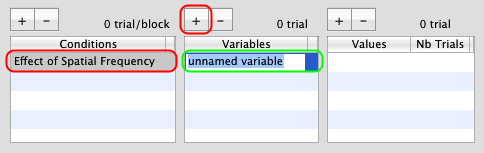
Make sure to select this new condition, and click on the '+' button to add a new variable (called unnamed variable by default). Rename this variable as sf (for spatial frequency).
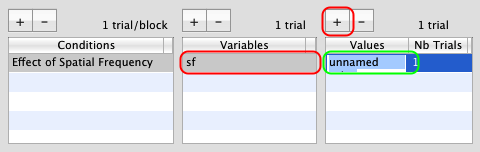
Make sure to select this new variable, and click on the '+' button to add a new value (called unnamed value by default). Set this value to 0.5. Leave the number of trials as 1.
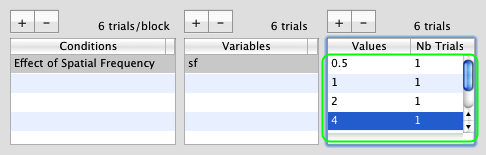
Add additional values for the sf variable, for example: 1, 2, 4, 8, and 16 (spatial frequency expressed in cycle per degree).
You have defined six experimental conditions: the sf variable is the independent variable of this experimental design which will take one of the six values in each trial. The number of trials for each value was left as 1 because the actual number is unknown since it will depend on how fast the adaptive method (staircase) can estimate the threshold.
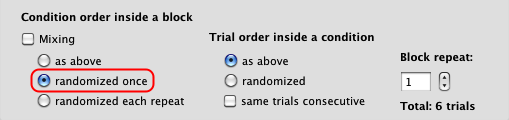
Finally, make sure to indicate that the trials order should be randomized.
Click on the OK button to validate the changes and return to the Designer panel.
Step 3: Updating the Hierarchy and the Stimulus Properties
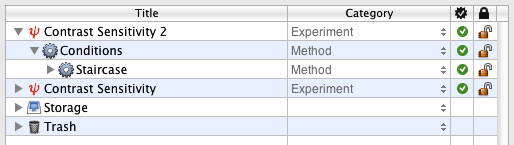
Click on and drag the Staircase event INTO the Conditions event so it is indented as illustrated. This way a staircase will run for each experimental condition defined above, and the different staircases will be presented interleaved.
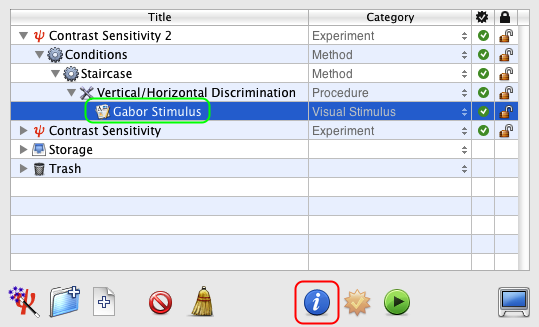
Option-click on the arrow in front of the Staircase event to reveal the entire hierarchy down to the Gabor Stimulus event.
Select the Gabor event and edit its properties by clicking on the Inspect button (or press Apple-i).
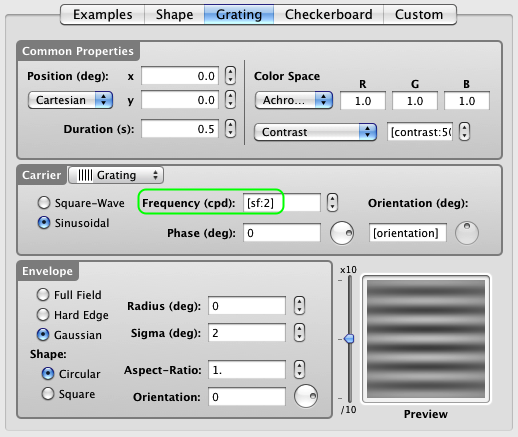
Finally, connect the sf variable, defined in the Conditions event, to the spatial frequency parameter of the stimulus (select and control-click on the text field content and choose the appropriate variable in the contextual menu). Note that the orientation variable has been already connected to the stimulus orientation in Lesson 1.
If you wish to preview the stimulus with a realistic value for its sf parameter (it is set to 0 by default), just add a colon (:) after the variable name followed by a value, all between brackets as illustrated. This will not affect the normal operation of the experiment in any way.
Click on the OK button to validate the changes and return to the Designer panel.
Check & run the Experiment!
Important note: To save the session results in the results database for future retrieval, always make sure to:
- run the experiment in full-screen mode by toggling the display icon in the Designer bottom toolbar,
- select the subject and group in the Session Information panel,
- click on the Run Session button in the Session Information panel.
Step 4: Visualizing the Results Summary
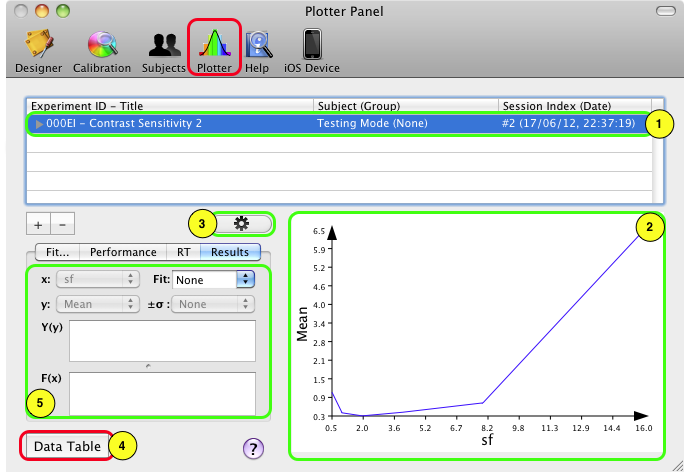
Upon termination of an experimental session, results data can be visualized immediately on the Plotter panel. Click on the Plotter icon in the toolbar to access this panel. The results for the last session appear at the top of the table.
As noted above the results are saved in the Psykinematix database only if the session was run in real mode. In this example, the Subject (Group) column shows Testing Mode (None) which indicates that the session has not been run in real mode and that its results will not be saved in the database after quitting Psykinematix.
To visualize a summary of the results:
1) Select the session,
2) If the measurements can be plotted as function of an independent variable, the graph area is automatically updated with default labels for the x- and y-axes,
3) The graph appearance can be customized by clicking on the properties button (through the graph properties palette), for example like:
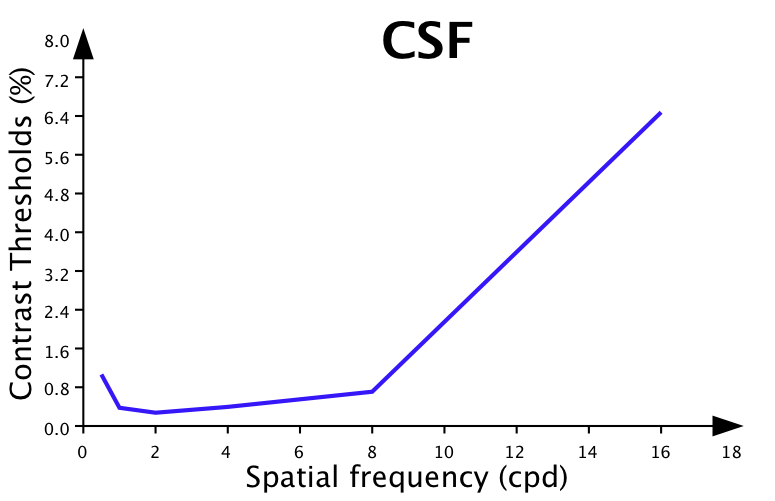
You can then print, copy, or save the plot to a file by control-clicking inside the graph area. You may also create a graph style for future use if you wish to create similar graphs for different data sets.
4) Click on the Data Table button to reveal the spreadsheet containing the results summary.
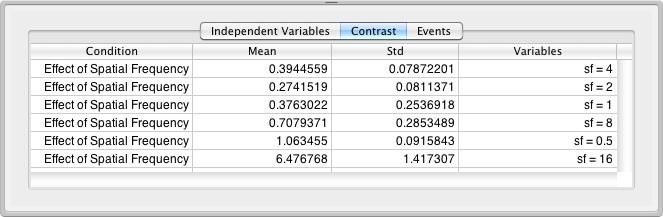
This table contains a single dataset named Contrast. This dataset presents the contrast threshold measured by the staircase method for each experimental condition. Each line refers to a condition, and each column presents the values for each relevant information:
- Condition: the name of the condition. Here a single condition was declared in the Conditions event (see Step 2 above).
- Mean and Std: the mean and standard deviation for the threshold measured by the staircase. The unit is the same as for the initial contrast value indicated in the Staircase event (see Step 6 in Lesson 1).
- Variables: the independent variables associated with the condition and their values. Several variables could be indicated in this column (see for example Lesson 2 of the Orientation Discrimination tutorial).
Control-click inside the spreadsheet to print or export this summary data.
For example, you could export the data directly to Excel to create a graphical representation of the contrast sensitivity that better suits your needs (eg. as the reciprocal of the contrast threshold as function of the spatial frequency) or to fit the data to a model using a dedicated data analysis software.
5) However instead of using a 3rd party software, you could also fit a CSF model to these data right from the Psykinematix software using the tools available on the left of the graph area.
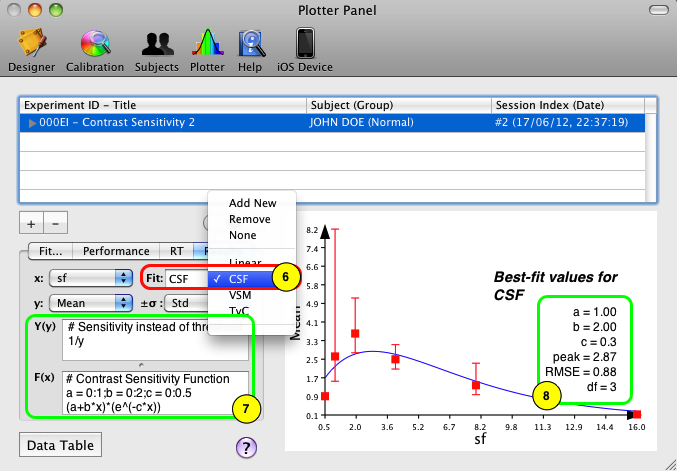
6) The simplest way to fit the data to a contrast sensitivity function (CSF) is to select the CSF built-in model in the Fit popup menu under the Results tab,
7) The analytical description of the selected model is presented in the Y(y) and F(x) text fields,
8) The result of the fitting procedure is immediately shown in the graph area with the plotted fitting function and its estimated parameters (to learn more about fitting, see the Data Fitting chapter).
Generally, there is no need to inspect further the results produced during the session. However, in case of doubts regarding the data reliability it is possible to review the dataset for each experimental condition as illustrated in the next step.
Step 5: Inspecting the Results for each Condition
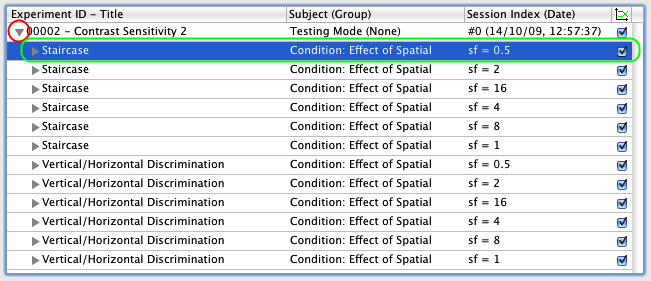
1) Click on the small arrow in front of the session entry to expand its second hierarchy level. You are presented with the method and procedure components of the results for each experimental condition: the first column indicates which method or procedure event produced the dataset, the second column indicates the condition, and the third column indicates the values of the independent parameters for this condition.
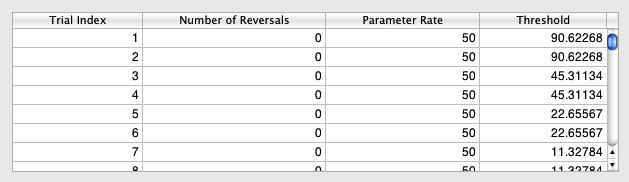
Selecting any Staircase entry updates the spreadsheet (revealed by clicking on the "Data Table" button) with the method parameters or variables for each trial run for the condition: the trial index, the number of reversals, the parameter rate and the value of the dependent variable (contrast Threshold).
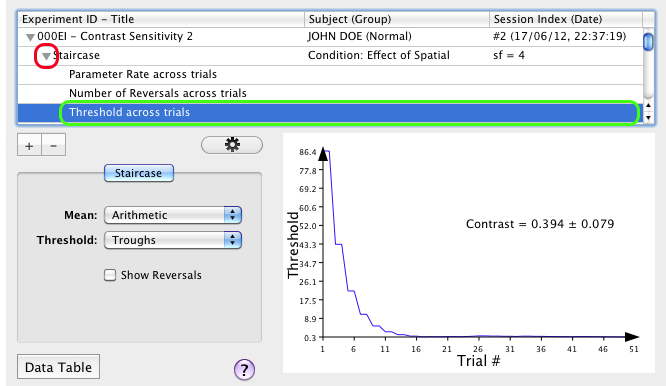
2) Click on the small arrow in front of the Staircase entry to expand the third hierarchy level. You are presented with the individual parameters or variables for the method. Select one of these entries to graph its attached dataset and update the spreadsheet with the trial history for the selected parameter or variable.
3) Click on the graph properties option to customize the graph appearance (through the graph properties palette), and control-click inside the graph to print, copy, or save it to a file.
See the "Import, Plot, Fit, and Export Data" tutorial to learn more about the Plotter panel.
Conclusion
In this lesson, you learned how to duplicate an experiment, specify experimental conditions through a Conditions method, move the events using drag and drop, and fit a model to measurements. You also learned how to inspect each level of the results hierarchy. Most important, you learned in Lessons 1 and 2 how to create a complete experiment to measure contrast sensitivity, one of the most fundamental properties of the visual system.
If you wish to expand this experiment by adding temporal masking or spatial context to the stimulus, or by using dynamic stimuli, you should consider the following lessons of the Orientation Discrimination tutorial to learn how to:
- implement a sandwich paradigm with forward and feedback masking noise (Lesson 3),
- add a spatial and temporal context (Lesson 4),
- add motion & dynamics through the use of a 1st-order drifting Gabor (Lesson 5).